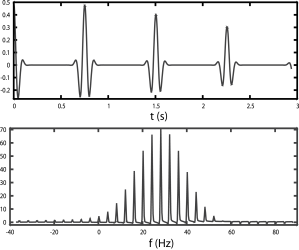
CPMG simulation/modeling
A script for simulating the CPMG sequence using the density matrix, with Gauss/Gauss envelope/spikelet broadening, for a forthcoming paper.
function [tot, gbb,t]=d_cpmg(N,tau,n,delta,r,rr)
%cpmg experiment evolution wjb 02/09
%simple/ideal pulse sequence:
%90y-n*[-180x-]
%N time steps
%tau
%n 180 loops
%delta chem shift
%r envelope gauss br^2
%rr spikelet gauss br^2
%matrix for I_x & I_y
a=[0 1/2; 1/2 0]; b=[0 -i/2; i/2 0];
%time step & initial rho
t=tau/(N-1); rho=a;
for k=1:n
sig(1)=trace(rho*a); sigi(1)=trace(rho*b);
for j=2:N/2
%iterate; free precession for tau/2
rho = [exp(-i*t*delta) 0; 0 exp(i*t*delta)]*rho*[exp(i*t*delta) 0; 0 exp(-i*t*delta)];
sig(j)=trace(rho*a); sigi(j)=trace(rho*b);
end
%apply 180x
rho = [0 exp(i*pi/2); exp(i*pi/2) 0]*rho*[0 exp(-i*pi/2); exp(-i*pi/2) 0];
%iterate; free precession for tau/2
sig(N/2+1)=trace(rho*a); sigi(N/2+1)=trace(rho*b);
for j=2:N/2
rho = [exp(-i*t*delta) 0; 0 exp(i*t*delta)]*rho*[exp(i*t*delta) 0; 0 exp(-i*t*delta)];
sig(j+N/2)=trace(rho*a); sigi(j+N/2)=trace(rho*b);
end
if k>1
tot=[tot (sig+i*sigi)];
else
tot=(sig+i*sigi);
end
end
tt=-tau/2:t:t*(N-1); gb=exp(-rr.*tt.^2); gb = [gb(N/2+1:N) gb(1:N/2)]; gbb=gb;
for i=1:n-1
gbb=[gbb gb];
end
t=0:t:t*(n*N-1); gbb=gbb.*exp(-r.*t.^2);